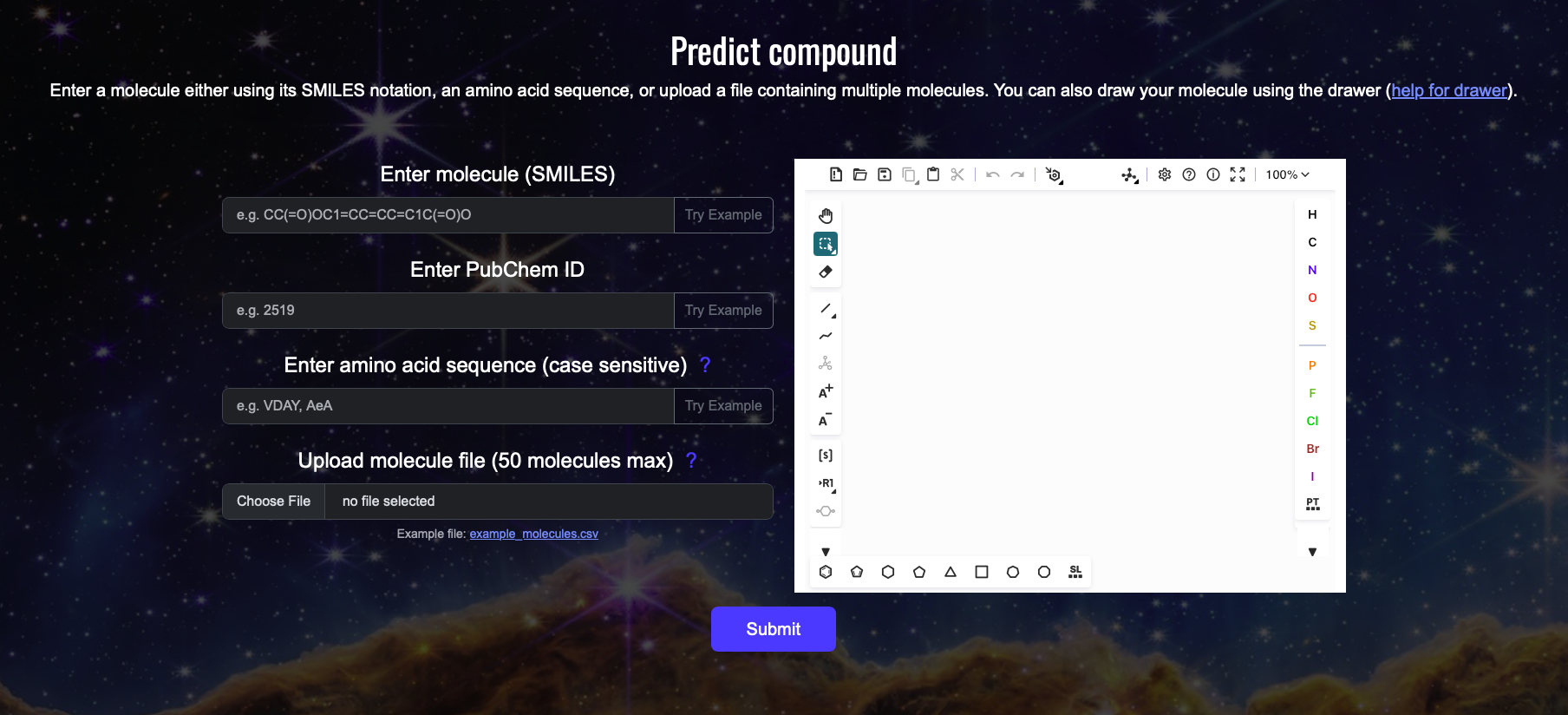
How to use the Predict page:
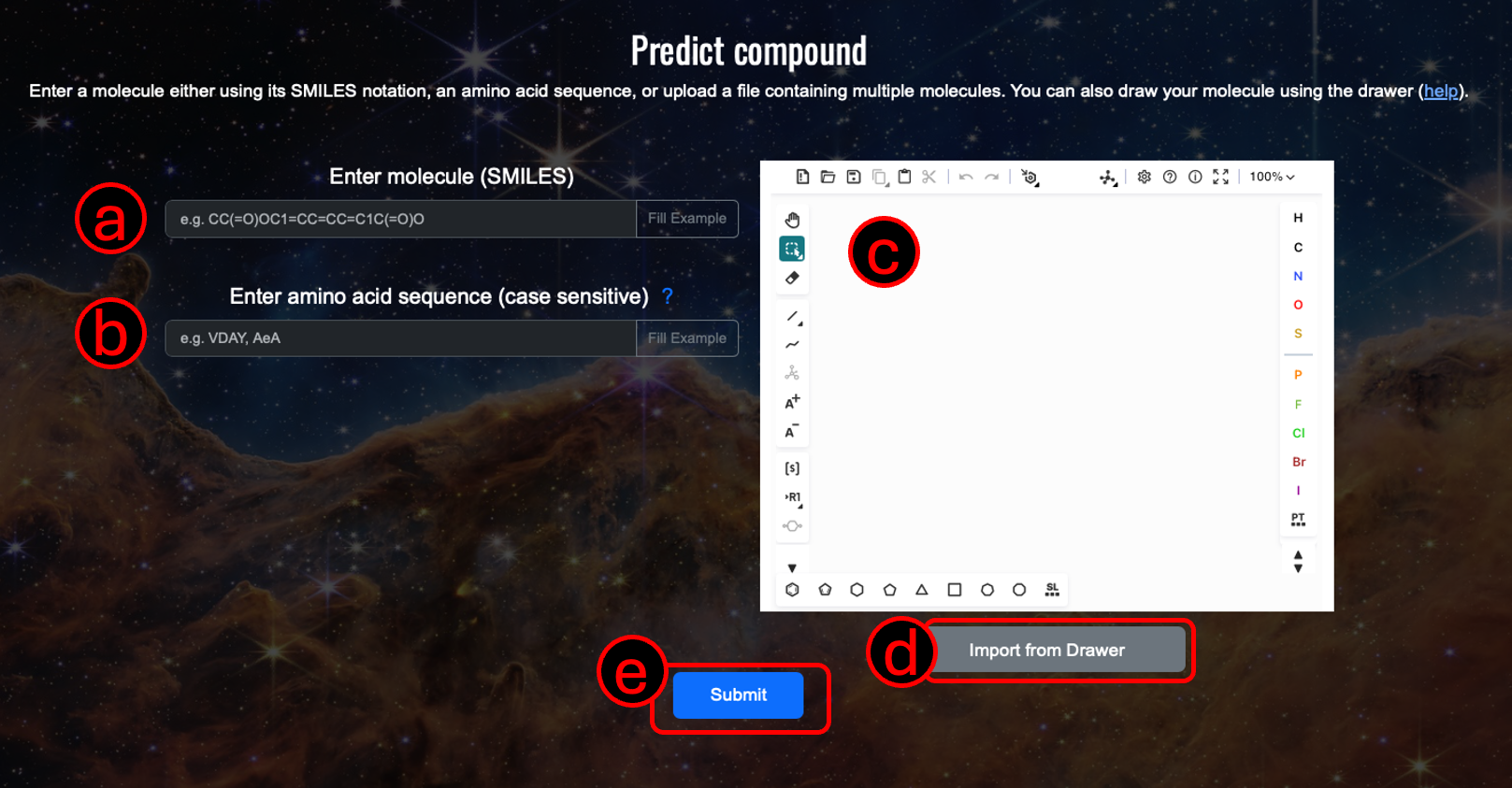
1. Input a molecule using (A) its SMILES notation, or (B) PubChem ID, or (C) amino acid sequence, or (D) upload a file containing the molecule information (see example file for reference, currently only .csv (comma seperated) file is supported, the file must contain two columns: value and type, type must be 'smiles' for now but we are expaning it to PubChem IDs and amino acid sequence).
IMPORTANT: The amino acid sequence field is case sensitive, use single-letter amino acid codes, UPPERCASE for L-amino acid and lowercase for D-amino acid, you can also mix L- and D-amino acids (e.g., AAeEEG).
Optional: Use (E) the molecule drawer to draw your molecule, check https://github.com/epam/ketcher/blob/master/documentation/help.md for guide.
2. Click the "Submit" button (F) to start the prediction process.
3. You will be redirected to the wait page, a unique link will be generated for your job submission. Wait for the results to be generated and get redirected to the results page.
Loading page
Loading page can return either (a) a waiting page if we are still processing your result, or (b) if the compound or file you submitted has errors, try submitting the example SMILES/amino acid/file/PubChem ID to check if the error is due to the input format or due to a server issue. If you are still having issues please contact us directly at: plete2026 [at] gmail.com

How to interpret the Results page:
The results page displays the predicted taste profiles for the submitted molecule. Each taste category (e.g., sweet, bitter, umami) is represented with a probability score ranging from 0%-100%.
In the page you can find:
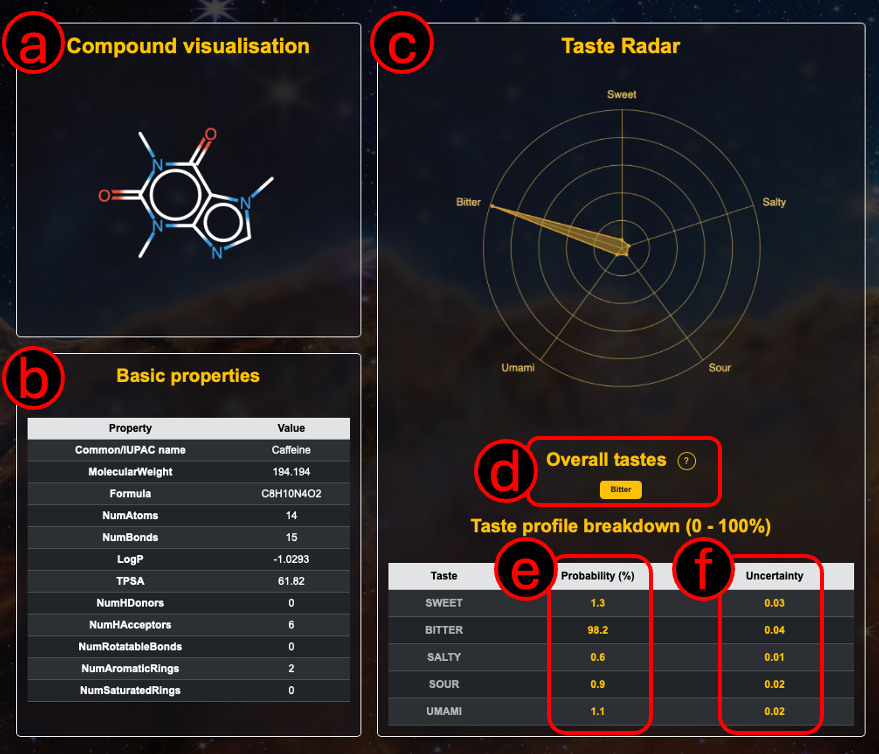
A. The visualised molecule (note that we use RDKit to canonicalise the input molecule, so the 2D structure may differ from your input) but nevertheless it is still the same compound.
B. Basic properties of the molecule, such as molecular weight, heavy atom count, molar refractivity, and topological polar surface area (TPSA) etc are calculated and shown.
C. A radar chart showing the predicted probabilities for each of the five basic tastes is provided (a higher probability is better).
D. Tastes that are above the 50% threshold are indicated as the overall predicted taste.
E. Probabilities for each basic taste, this is the same data as is presented in (C) in the form of a radar chart, here it is in the form of a table.
F. Uncertainty quantification for each of the five basic taste predictions (ranging from 0-1; a lower uncertainty is better).
G. A download button that allows you to save the result as a CSV file, which includes the predicted probabilities and uncertainties for each of the five taste categories.
How to use the Multi-Compound Results page:
Multi-Compound result page only contains the download button, the download button allows you to download a CSV file containing the predicted probabilities and uncertainties for each of the five taste categories for all the compounds you submitted in your batch job.
Walkthrough Example
Follow this example which illustrates how to obtain the SMILES format for a compound from PubChem, how to modify the compound (in this case an N-demethylation is performed) and the modified molecule is then submitted to PLETE. We will also explain how to interpret the results obtained from the web server.
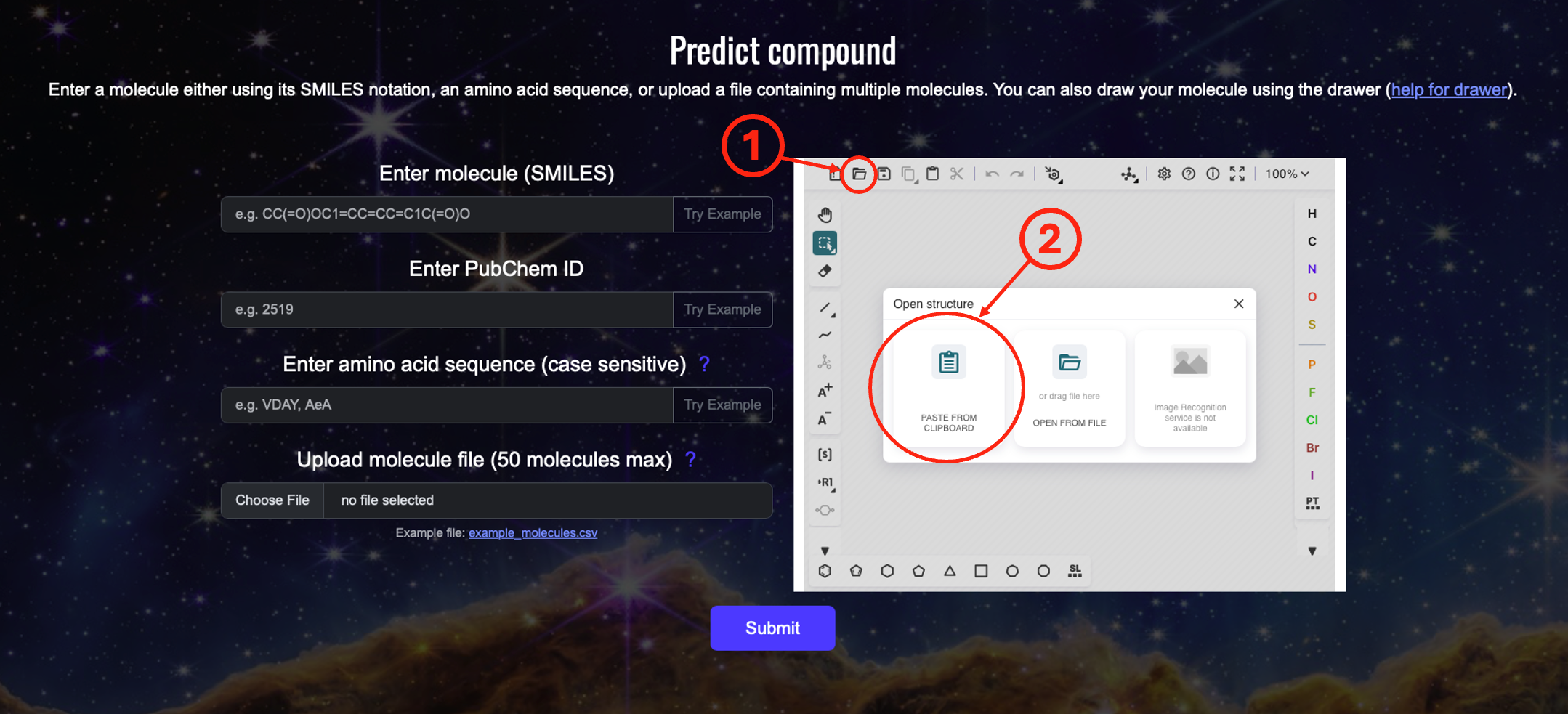
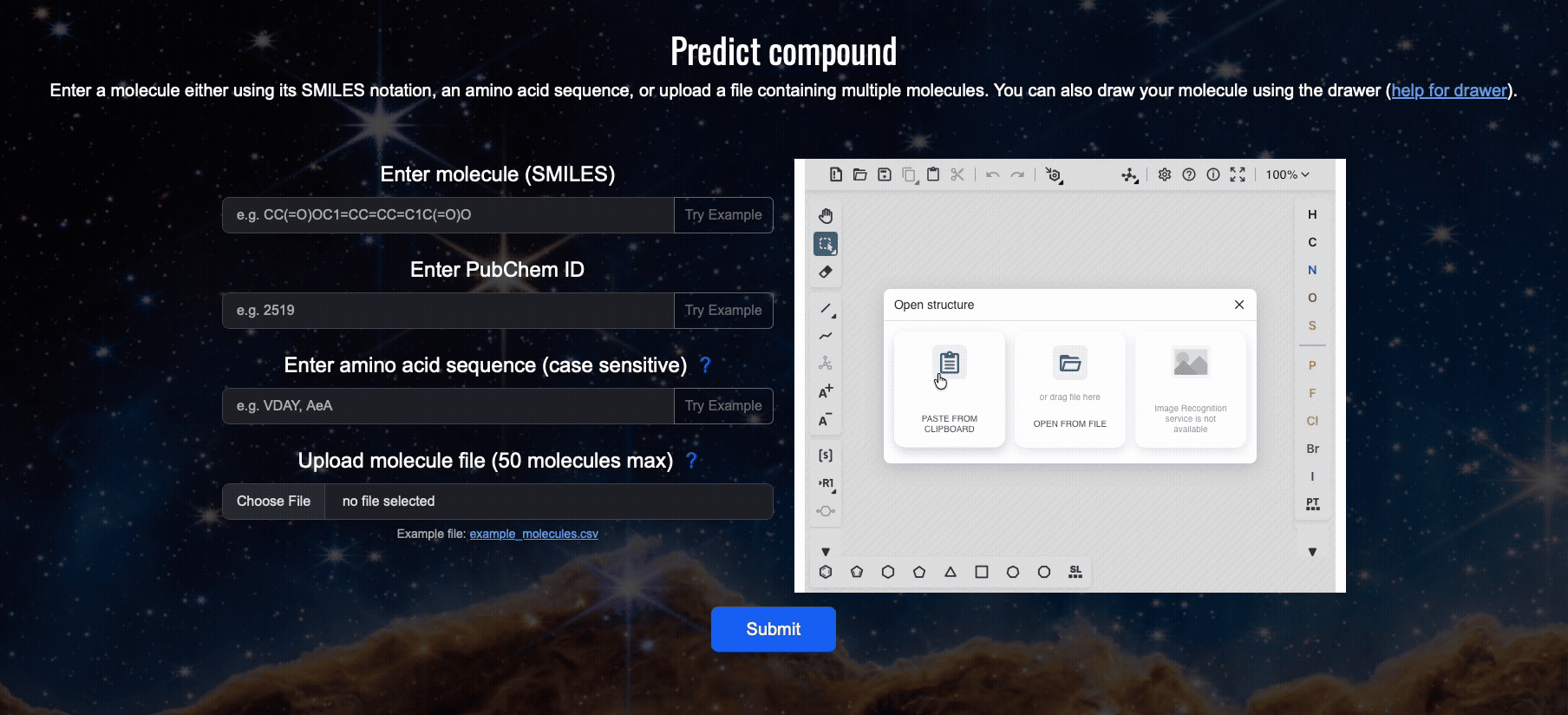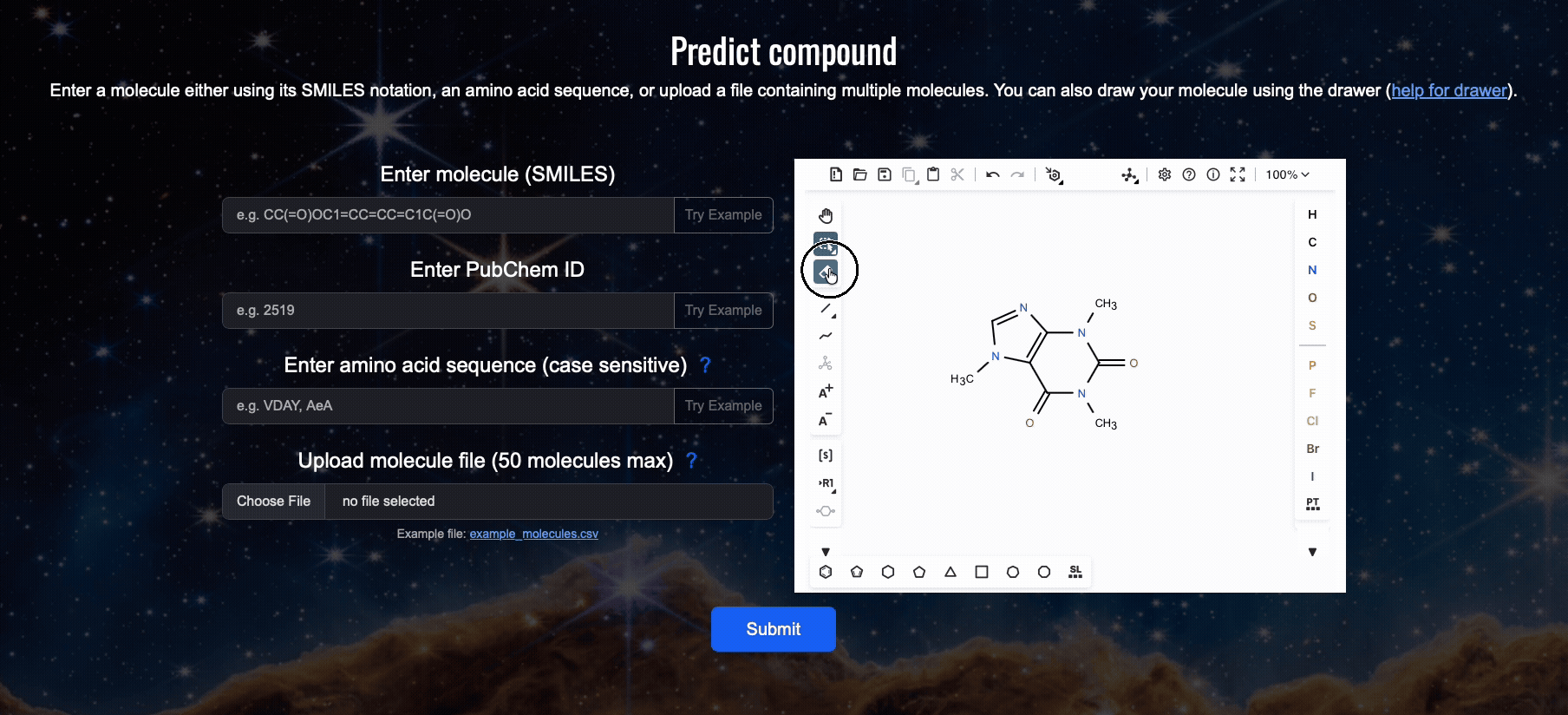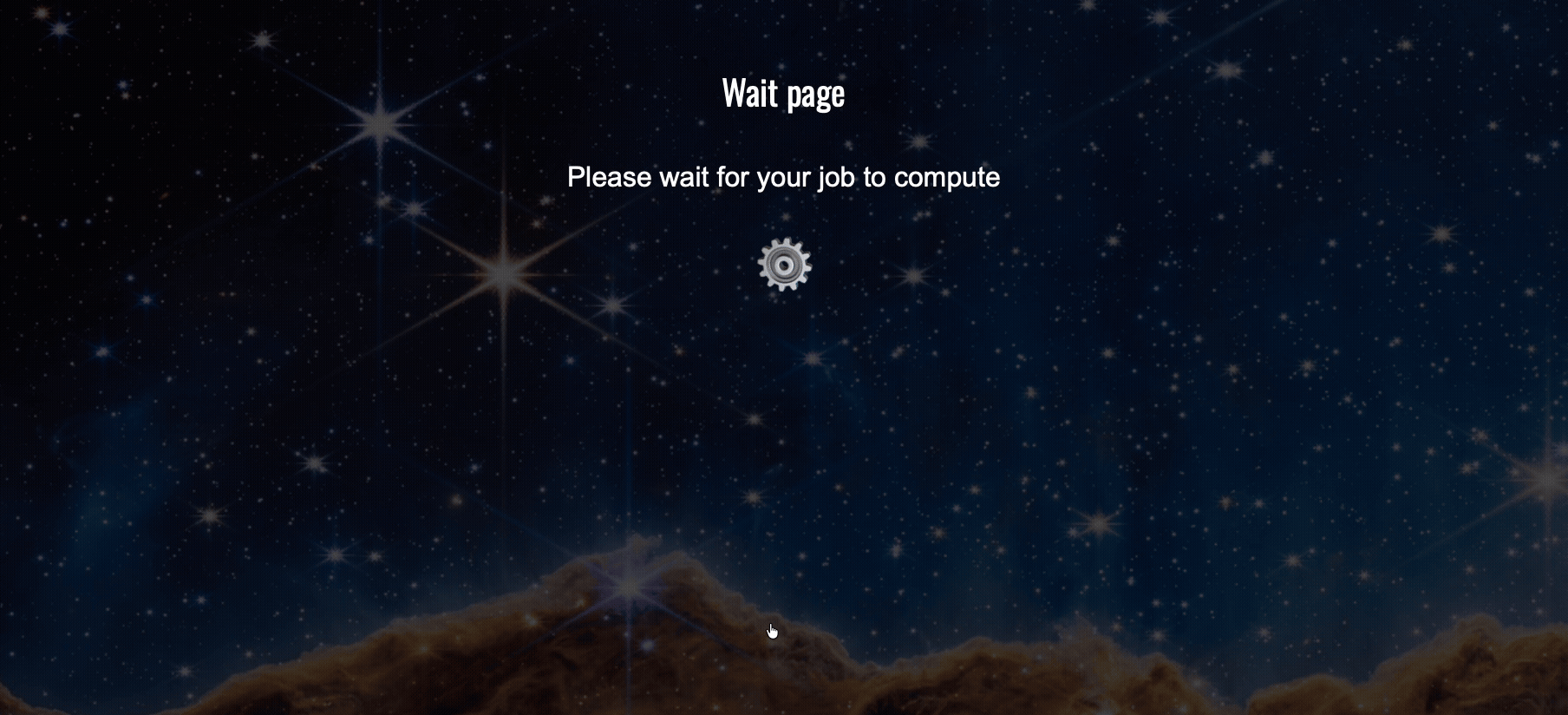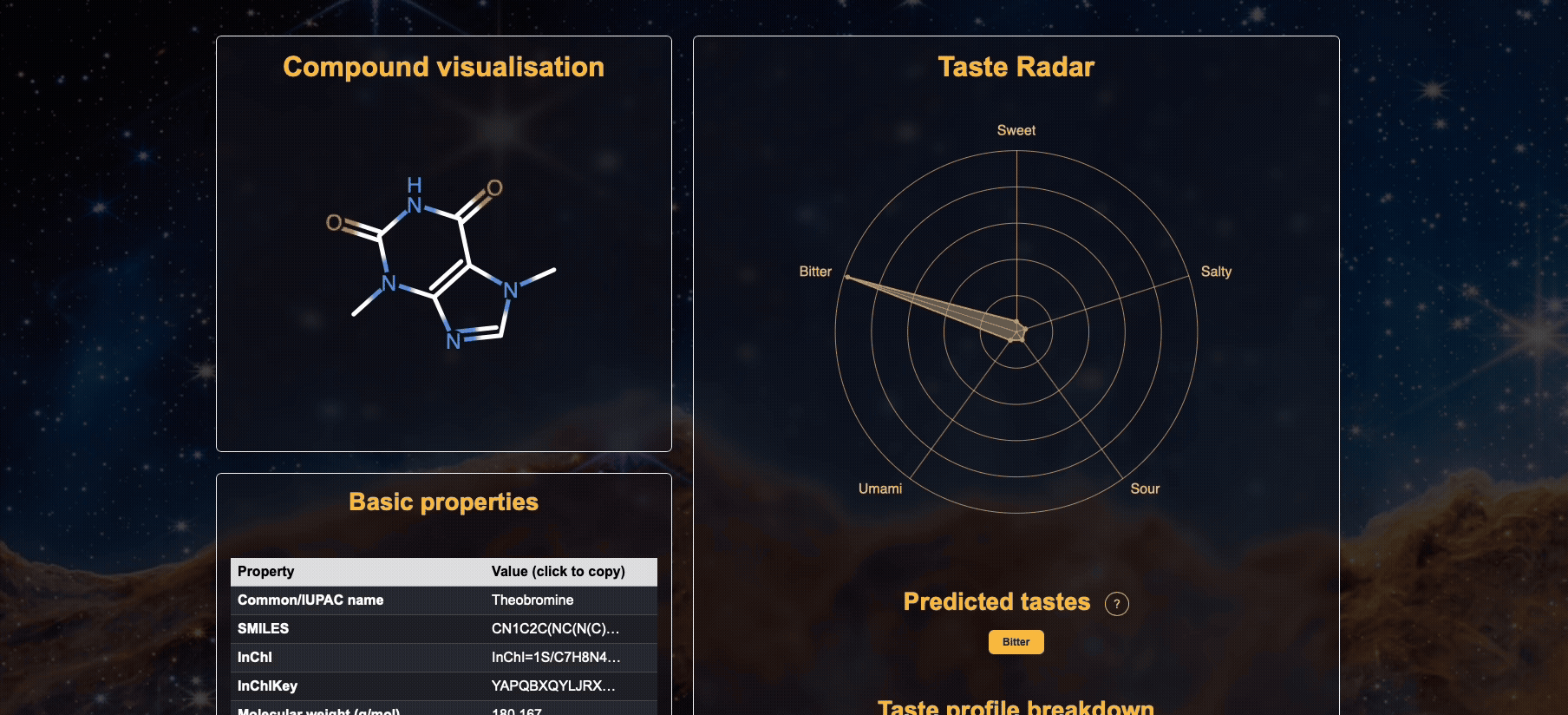
Step 2: Open the compound load window in the drawer
Click (1) and (2) "PASTE FROM CLIPBOARD". The drawer will open a window which you can later use to load a compound using its SMILES.
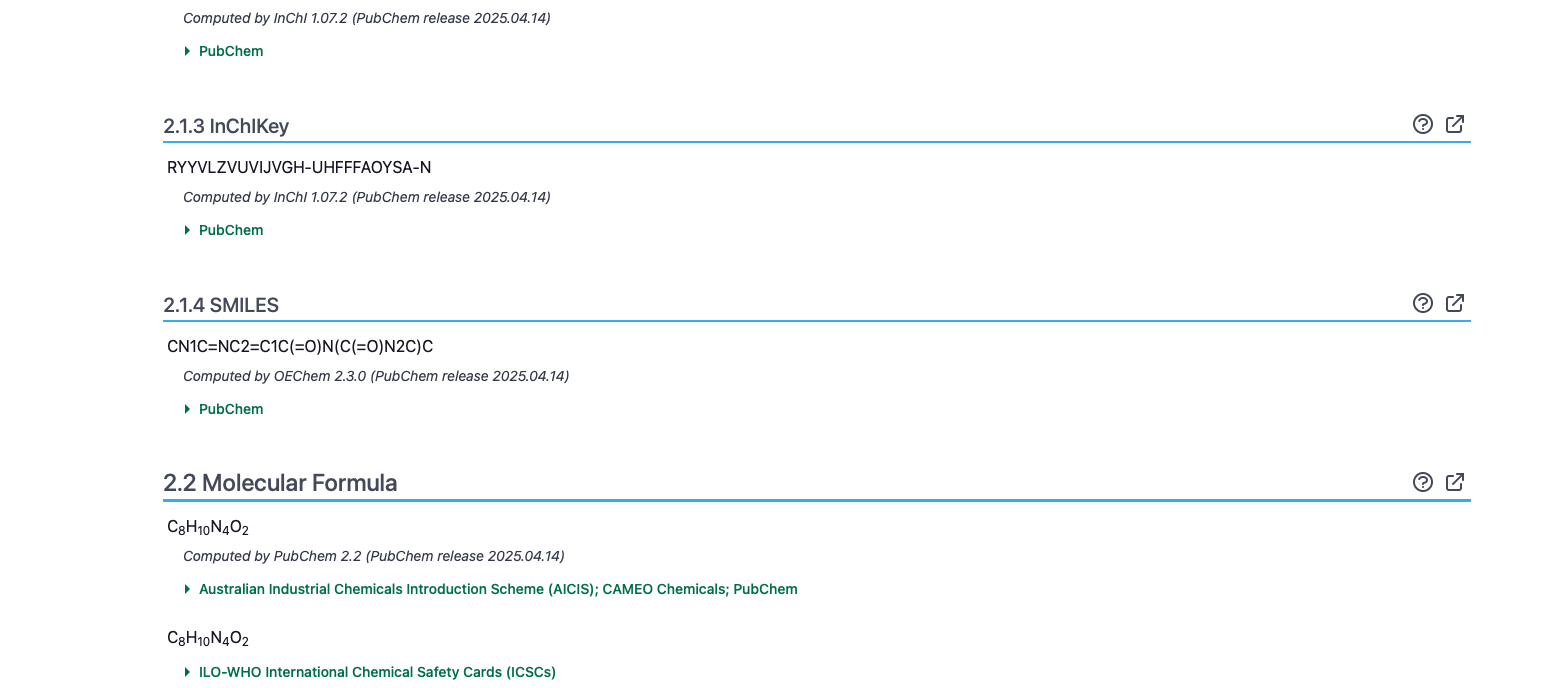
Step 3: Obtain the SMILES from PubChem website
For this example, we use caffeine. Whenever possible, get SMILES from PubChem, which includes stereochemistry information
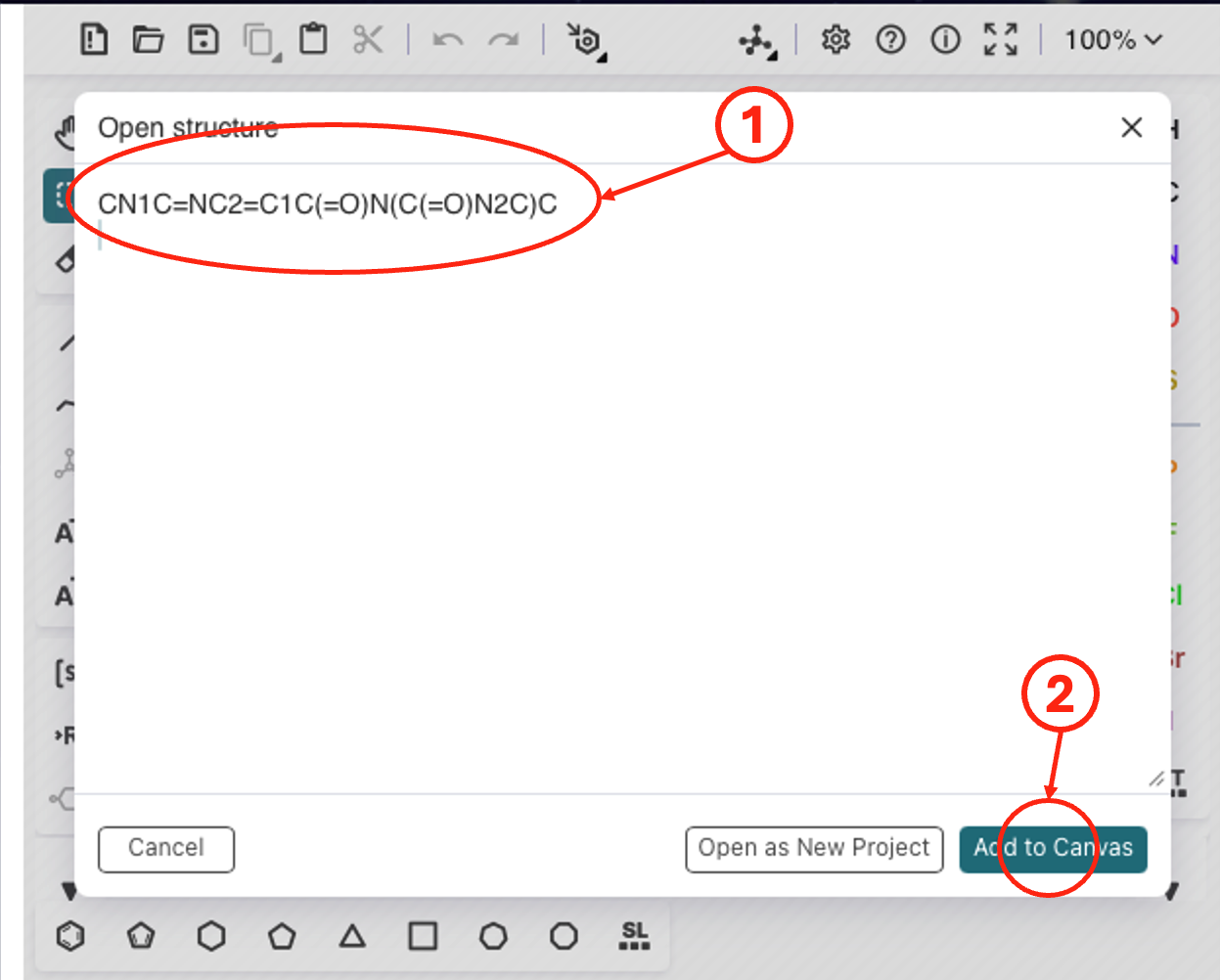
(1) Paste the SMILES, then (2) add the compound to the drawer window.
Step 4: Paste the SMILES from the previous step and add to the drawer canvas, modify the compound (e.g., N-demethylation) in the drawer. Load the modified SMILES and submit it for prediction.
(1) Paste the SMILES, then (2) add the compound to the drawer window.
For all web server related issues please contact Kenny Lam directly at: plete2026 [at] gmail.com